Note
This page was generated from a Jupyter Notebook. Download the notebook (.ipynb)
[1]:
# Skipped in CI: Colab/bootstrap dependency install cell.
Coupling Analysis: Multi-Channel Coupling
This tutorial demonstrates the Coupling Function (CF) analysis workflow using gwexpy.analysis.CouplingFunctionAnalysis.
What is a Coupling Function?
A coupling function quantifies how much a witness (environmental) sensor’s noise couples into a target (sensitive) channel. It is defined as:
where \(P_{\rm inj}\) and \(P_{\rm bkg}\) are the power spectral densities during injection and background periods.
Typical application: PEM (Physical Environment Monitor) injection analysis in gravitational-wave detectors.
Note — SigmaThreshold statistical validity: When the number of averaged segments
n_avgis below 10, the Gaussian approximation of the χ²(2K) distribution becomes less accurate. AUserWarningwill be raised in this case. Consider usingPercentileThresholdfor short data segments.
[2]:
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
import matplotlib.pyplot as plt
import numpy as np
from astropy import units as u
from gwexpy.analysis.coupling import (
CouplingFunctionAnalysis,
RatioThreshold,
SigmaThreshold,
)
from gwexpy.timeseries import TimeSeries, TimeSeriesDict
1. Prepare Synthetic Injection / Background Data
We simulate:
A witness channel (environmental sensor): seismic or acoustic
Two target channels (sensitive instruments): one strongly coupled, one weakly coupled
The injection adds a coherent tone to the witness. The targets respond proportionally to the coupling strength.
[3]:
rng = np.random.default_rng(0)
fs = 512 # sample rate [Hz]
T = 32.0 # duration [s]
n = int(fs * T)
t = np.arange(n) / fs
dt = (1 / fs) * u.s
# Frequency bins for injected tones
f_inj1 = 12.5 # Hz
f_inj2 = 25.0 # Hz
# Noise floor
noise_level = 0.1
# --- Background data (no injection) ---
wit_bkg = rng.normal(0, noise_level, n)
tgt1_bkg = rng.normal(0, noise_level, n)
tgt2_bkg = rng.normal(0, noise_level, n)
# --- Injection data: excite the witness with tones ---
inj_signal = 5.0 * np.sin(2 * np.pi * f_inj1 * t) + 3.0 * np.sin(2 * np.pi * f_inj2 * t)
wit_inj = inj_signal + rng.normal(0, noise_level, n)
# Target 1: strong coupling (CF ≈ 0.4 at f_inj1, 0.3 at f_inj2)
cf_true1 = 0.4
tgt1_inj = cf_true1 * inj_signal + rng.normal(0, noise_level, n)
# Target 2: weak coupling (CF ≈ 0.05)
cf_true2 = 0.05
tgt2_inj = cf_true2 * inj_signal + rng.normal(0, noise_level, n)
# Wrap in TimeSeries
def make_ts(data, name, unit=u.ct):
return TimeSeries(data, dt=dt, t0=0*u.s, unit=unit, name=name)
# Background dict
data_bkg = TimeSeriesDict({
"ACC:FLOOR_X": make_ts(wit_bkg, "ACC:FLOOR_X", u.m / u.s**2),
"GW:DARM": make_ts(tgt1_bkg, "GW:DARM", u.m),
"GW:AUX": make_ts(tgt2_bkg, "GW:AUX", u.m),
})
# Injection dict
data_inj = TimeSeriesDict({
"ACC:FLOOR_X": make_ts(wit_inj, "ACC:FLOOR_X", u.m / u.s**2),
"GW:DARM": make_ts(tgt1_inj, "GW:DARM", u.m),
"GW:AUX": make_ts(tgt2_inj, "GW:AUX", u.m),
})
print("Background channels:", list(data_bkg.keys()))
print("Injection channels: ", list(data_inj.keys()))
Background channels: ['ACC:FLOOR_X', 'GW:DARM', 'GW:AUX']
Injection channels: ['ACC:FLOOR_X', 'GW:DARM', 'GW:AUX']
2. Run Coupling Function Analysis
CouplingFunctionAnalysis.compute() takes the injection and background TimeSeriesDict, and the name of the witness channel.
[4]:
cfa = CouplingFunctionAnalysis()
results = cfa.compute(
data_inj=data_inj,
data_bkg=data_bkg,
fftlength=4.0, # 4-second FFT segments
overlap=0.5, # 50% overlap
witness="ACC:FLOOR_X", # the environmental sensor
threshold_witness=RatioThreshold(ratio=10.0), # witness must show 10× excess
threshold_target=RatioThreshold(ratio=2.0), # target must show 2× excess
)
print("Results computed for:", list(results.keys()))
Results computed for: ['GW:DARM', 'GW:AUX']
3. Inspect Results
Each result is a CouplingResult containing the coupling function, upper limit, and diagnostic PSDs.
[5]:
# Strong coupling channel (GW:DARM)
res_darm = results["GW:DARM"]
print("CF valid bins:", res_darm.valid_mask.sum())
print("CF at f_inj1 bin:", end=" ")
freqs = res_darm.cf.frequencies.value
idx1 = np.argmin(np.abs(freqs - f_inj1))
idx2 = np.argmin(np.abs(freqs - f_inj2))
print(f"{res_darm.cf.value[idx1]:.3f} (expected ~{cf_true1})")
print(f"CF at f_inj2 bin: {res_darm.cf.value[idx2]:.3f} (expected ~{cf_true1})")
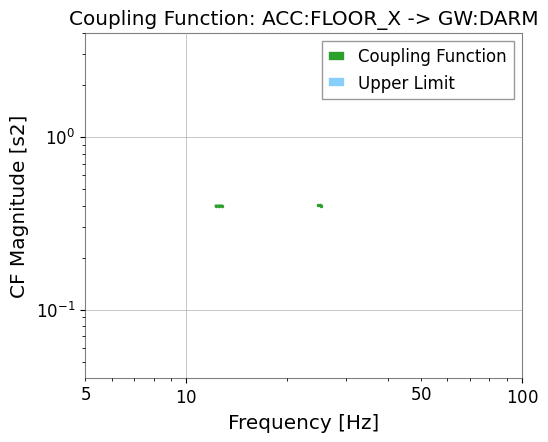
# Simple CF plot
res_darm.plot_cf(xlim=(5, 100));
CF valid bins: 6
CF at f_inj1 bin: 0.400 (expected ~0.4)
CF at f_inj2 bin: 0.401 (expected ~0.4)
[6]:
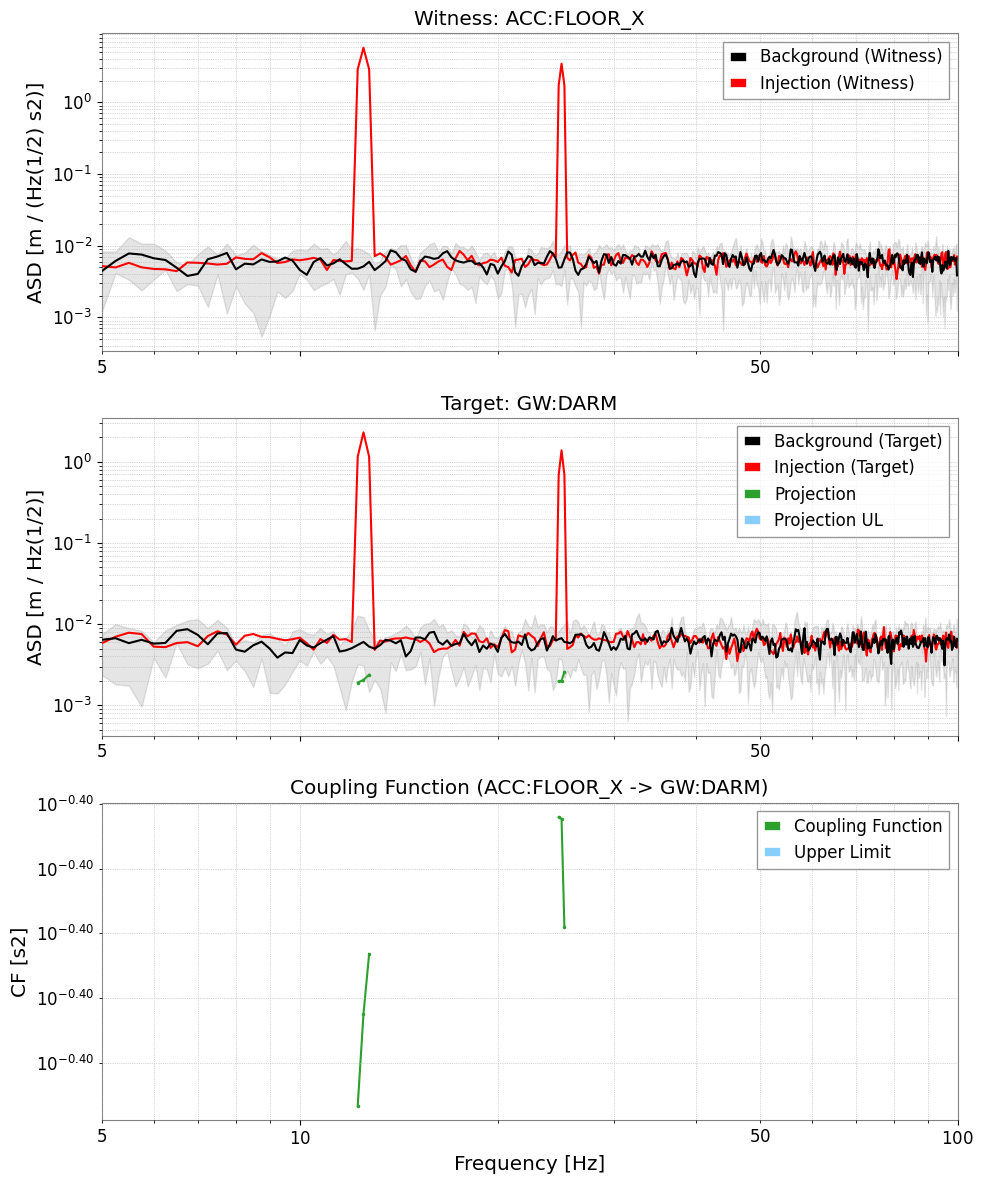
# Full diagnostic plot: Witness ASD | Target ASD | CF
res_darm.plot(xlim=(5, 100));
[7]:
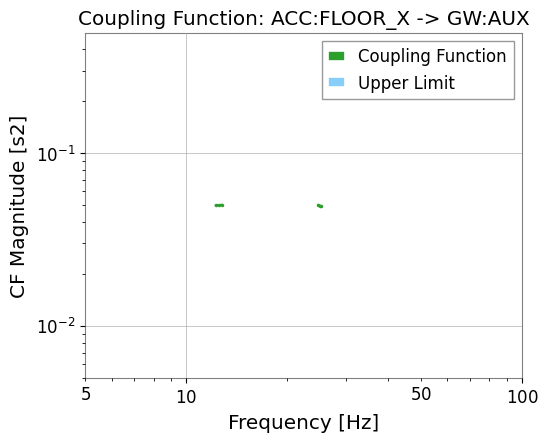
# Weak coupling channel (GW:AUX)
res_aux = results["GW:AUX"]
print("GW:AUX valid CF bins:", res_aux.valid_mask.sum())
res_aux.plot_cf(xlim=(5, 100));
GW:AUX valid CF bins: 6
import warnings
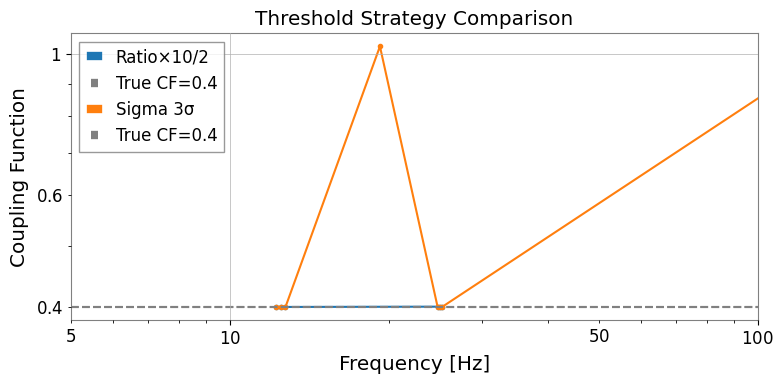
4. Threshold Strategies
gwexpy provides three built-in threshold strategies:
Strategy |
Description |
Best for |
|---|---|---|
|
\(P_{\rm inj} > r \cdot P_{\rm bkg}\) |
Fast screening, physical reasoning |
|
Gaussian significance test |
Stationary, Gaussian backgrounds |
|
Empirical distribution |
Non-Gaussian, glitchy backgrounds |
[8]:
import warnings
with warnings.catch_warnings():
warnings.simplefilter('ignore')
# Compare threshold strategies
n_avg = T / 4.0 # approximate averaging count for 4-second FFT
strategies = {
"Ratio×10/2": (RatioThreshold(10.0), RatioThreshold(2.0)),
"Sigma 3σ": (SigmaThreshold(3.0), SigmaThreshold(2.0)),
}
fig, ax = plt.subplots(figsize=(8, 4))
for label, (thr_w, thr_t) in strategies.items():
res = cfa.compute(
data_inj=data_inj,
data_bkg=data_bkg,
fftlength=4.0,
overlap=0.5,
witness="ACC:FLOOR_X",
threshold_witness=thr_w,
threshold_target=thr_t,
)
cf = res["GW:DARM"].cf
valid = ~np.isnan(cf.value)
ax.plot(cf.frequencies.value[valid], cf.value[valid], marker=".", label=label)
ax.axhline(cf_true1, color="gray", linestyle="--", label=f"True CF={cf_true1}")
ax.set_xscale("log")
ax.set_yscale("log")
ax.set_xlim(5, 100)
ax.set_xlabel("Frequency [Hz]")
ax.set_ylabel("Coupling Function")
ax.set_title("Threshold Strategy Comparison")
ax.legend()
plt.tight_layout()
5. Frequency Range Restriction
Use frange to restrict the coupling function evaluation to a specific frequency band.
[9]:
results_band = cfa.compute(
data_inj=data_inj,
data_bkg=data_bkg,
fftlength=4.0,
overlap=0.5,
witness="ACC:FLOOR_X",
frange=(10.0, 50.0), # only evaluate 10–50 Hz
)
cf_band = results_band["GW:DARM"].cf
valid_band = ~np.isnan(cf_band.value)
print(f"Valid CF bins in 10–50 Hz: {valid_band.sum()}")
results_band["GW:DARM"].plot_cf(xlim=(5, 100));
Valid CF bins in 10–50 Hz: 6
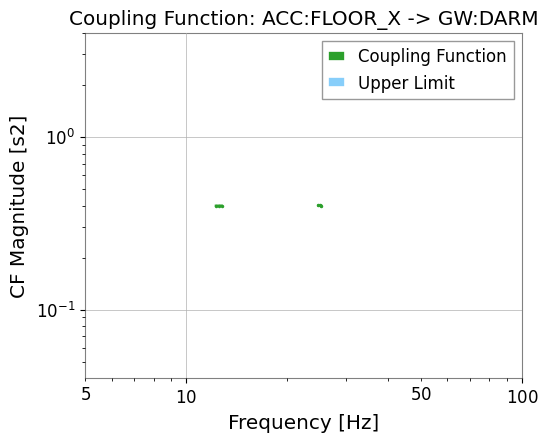
6. Upper Limits
At frequencies where the witness is excited but the target shows no measurable excess, gwexpy computes an upper limit on the coupling function.
This is useful for setting constraints even when the coupling is below the noise floor.
[10]:
# Upper limit is automatically computed in cf_ul attribute
res_darm = results["GW:DARM"]
cf_ul = res_darm.cf_ul
ul_valid = ~np.isnan(cf_ul.value)
print(f"Upper limit bins: {ul_valid.sum()}")
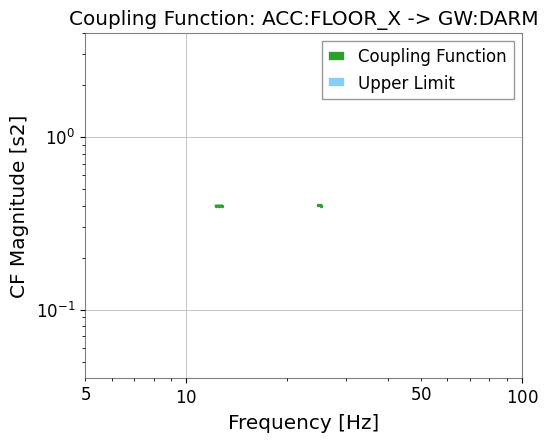
# The full diagnostic plot includes both CF and upper limit
res_darm.plot_cf(xlim=(5, 100));
Upper limit bins: 0
Summary
from gwexpy.analysis.coupling import CouplingFunctionAnalysis, RatioThreshold
cfa = CouplingFunctionAnalysis()
results = cfa.compute(
data_inj=data_inj, # TimeSeriesDict with injection data
data_bkg=data_bkg, # TimeSeriesDict with background data
fftlength=4.0,
witness="ACC:FLOOR_X",
threshold_witness=RatioThreshold(25.0),
threshold_target=RatioThreshold(4.0),
)
# Access result for one target
res = results["GW:DARM"]
res.plot() # diagnostic 3-panel plot
res.cf # FrequencySeries: coupling function values
res.cf_ul # FrequencySeries: upper limits
See also: