Note
This page was generated from a Jupyter Notebook. Download the notebook (.ipynb)
[1]:
# Skipped in CI: Colab/bootstrap dependency install cell.
Histogram: Basics
This tutorial introduces gwexpy.histogram.Histogram and demonstrates:
Creating histograms (from totals, from events using
.fill())Visualization (counts and density)
Basic statistics (mean, median, variance, quantiles)
Rebinning with covariance propagation
Integration with error propagation
Serialization (HDF5 and JSON)
ROOT interoperability (optional; handled with
try/except)
Run this notebook in the gwexpy conda environment so that all dependencies are available.
[2]:
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
# Environment check
import sys
import matplotlib.pyplot as plt
import numpy as np
from astropy import units as u
from gwexpy.histogram import Histogram
print('python', sys.version.splitlines()[0])
print('numpy', np.__version__)
print('astropy', u.__version__ if hasattr(u, '__version__') else 'unknown')
python 3.11.15 | packaged by conda-forge | (main, Mar 5 2026, 16:45:40) [GCC 14.3.0]
numpy 2.4.4
astropy unknown
1. Creating Histograms
Two common ways to create a Histogram:
Directly from bin totals and edges
From raw events using
.fill()(supportsastropy.units.Quantityweights)
[3]:
# 1.1 Direct construction from totals
edges = np.array([0.0, 1.0, 2.0, 4.0])
values = np.array([10.0, 25.0, 15.0]) # bin totals
h = Histogram(values=values, edges=edges, unit='count')
h
[3]:
<Histogram (nbins=3, unit=ct)>
[4]:
# 1.2 Creating from events with weights (Quantity)
rng = np.random.RandomState(42)
events = np.concatenate([
rng.normal(0.5, 0.1, size=50),
rng.normal(1.6, 0.2, size=120),
rng.normal(3.0, 0.3, size=80)
])
weights = u.Quantity(rng.uniform(0.5, 2.0, size=events.size), unit='count')
h2 = Histogram(values=np.zeros(len(edges)-1), edges=edges, unit='count').fill(events, weights=weights)
h2
[4]:
<Histogram (nbins=3, unit=ct)>
2. Visualization
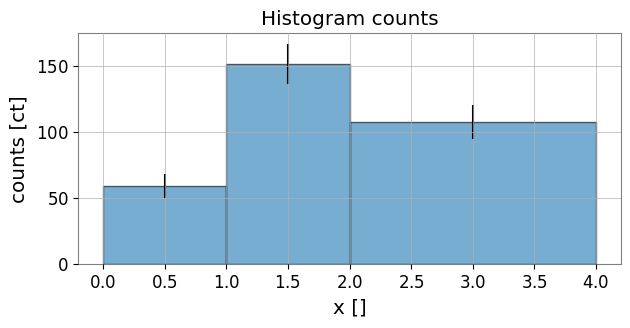
Plot bin totals (with errorbars if available) and density (value / bin width).
[5]:
# Plot counts with errorbars
fig, ax = plt.subplots(figsize=(7,3))
centers = h2.bin_centers.value
widths = h2.bin_widths.value
counts = h2.values.value
errs = (h2.errors.value if h2.errors is not None else None)
ax.bar(centers, counts, width=widths, align='center', edgecolor='k', alpha=0.6)
if errs is not None:
ax.errorbar(centers, counts, yerr=errs, fmt='none', ecolor='k')
ax.set_xlabel(f"x [{h2.xunit}]")
ax.set_ylabel(f"counts [{h2.unit}]")
ax.set_title('Histogram counts')
plt.show()
[6]:
# Density plot (values / bin width)
density = h2.to_density()
fig, ax = plt.subplots(figsize=(7,2))
ax.step(h2.edges.value, np.append(density.value, density.value[-1]), where='post')
ax.set_xlabel(f"x [{h2.xunit}]")
ax.set_ylabel(f"density [{density.unit}]")
ax.set_title('Histogram density (values / bin width)')
plt.show()
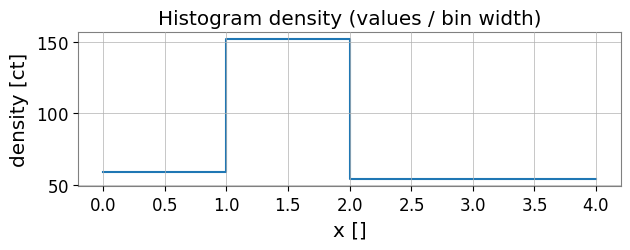
3. Basic statistics
Compute mean, median, variance and quantiles. quantile() is robust to flat bins.
[7]:
print('mean:', h2.mean())
print('median:', h2.median())
print('variance:', h2.var())
print('std:', h2.std())
print('quantile(0.25):', h2.quantile(0.25))
mean: 1.8213054429831148
median: 1.6599075323583463
variance: 0.8432605001074853
std: 0.918292164894967
quantile(0.25): 1.1348114645471883
4. Rebinning
Demonstrate rebinning to non-uniform bins and verify totals are conserved and covariance is propagated.
[8]:
new_edges = np.array([0.0, 1.5, 4.0])
h_reb = h2.rebin(new_edges)
print('sum original:', h2.values.sum())
print('sum rebinned:', h_reb.values.sum())
print('cov present original:', h2.cov is not None)
print('cov shape rebinned:', h_reb.cov.shape if h_reb.cov is not None else None)
sum original: 318.85040365900875 ct
sum rebinned: 318.85040365900875 ct
cov present original: False
cov shape rebinned: None
5. Integration with error propagation
Integrate over a partial interval (handles partial bin overlap) and show the propagated error.
[9]:
I = h2.integral(0.7, 2.2)
I_val, I_err = h2.integral(0.7, 2.2, return_error=True)
print('Integral (no error):', I)
print('Integral with error:', I_val, I_err)
Integral (no error): 180.3597019890495 ct
Integral with error: 180.3597019890495 ct 15.17057636890394 ct
6. Serialization (HDF5 and JSON)
Show HDF5 write/read and a JSON-friendly dict for light-weight exchange.
[10]:
# HDF5 roundtrip
import tempfile
from pathlib import Path
hist_path = Path(tempfile.gettempdir()) / 'gwexpy_example_hist.h5'
h2.write(hist_path, format='hdf5', path='hist1')
h_loaded = Histogram.read(hist_path, format='hdf5', path='hist1')
print('HDF5 roundtrip equal (values):', np.allclose(h2.values.value, h_loaded.values.value))
HDF5 roundtrip equal (values): True
[11]:
import json
def hist_to_json_dict(h):
def get_val(x):
return x.value if hasattr(x, 'value') else x
return {
'values': h.values.value.tolist(),
'edges': h.edges.value.tolist(),
'unit': str(h.unit),
'xunit': str(h.xunit),
'cov': (h.cov.value.tolist() if h.cov is not None else None),
'sumw2': (h.sumw2.value.tolist() if h.sumw2 is not None else None),
'underflow': float(get_val(getattr(h, 'underflow', 0.0))),
'overflow': float(get_val(getattr(h, 'overflow', 0.0))),
}
def hist_from_json_dict(d):
kwargs = {}
if d.get('cov') is not None:
kwargs['cov'] = np.array(d['cov'])
if d.get('sumw2') is not None:
kwargs['sumw2'] = np.array(d['sumw2'])
if d.get('underflow') is not None:
kwargs['underflow'] = d['underflow']
if d.get('overflow') is not None:
kwargs['overflow'] = d['overflow']
return Histogram(values=np.array(d['values']), edges=np.array(d['edges']), unit=d['unit'], xunit=d['xunit'], **kwargs)
j = hist_to_json_dict(h2)
open('hist.json','w').write(json.dumps(j, indent=2))
h_j = hist_from_json_dict(json.loads(open('hist.json').read()))
print('JSON roundtrip (values):', np.allclose(h2.values.value, h_j.values.value))
JSON roundtrip (values): True
7. ROOT interoperability (optional)
PyROOT may not be present in all environments. Use try/except to handle absence gracefully.
[12]:
from gwexpy.interop.root_ import from_root, to_th1d
try:
th1 = to_th1d(h2)
print('TH1D created:', th1.GetName(), 'nbins:', th1.GetNbinsX())
print('ROOT underflow:', th1.GetBinContent(0))
# Optionally convert back if supported
try:
h_from_root = from_root(th1) # use from_root from the module
print('Back to Histogram:', h_from_root)
except Exception as e:
print('Round-trip from ROOT not supported or error:', e)
except Exception as e:
print('ROOT not available or error:', e)
ROOT not available or error: The 'ROOT' package is required for this feature but is not installed. You can install it via 'pip install ROOT'.
8. Exercises
Generate a 1D mixture of Gaussians and verify rebin totals are preserved.
Create weighted events where weights are
Quantityand verifysumw2updates assum(w^2).Save a histogram to HDF5 and JSON, and load it back in a separate Python process to check equality.
Summary
Histogramstores bin totals and propagates covariance under linear transforms (rebin).fill()supportsQuantityweights and updatessumw2/covappropriately.Use HDF5 for lossless IO; JSON for light-weight exchange.
For very large
nbins, consider sparse/low-rank covariance representations.