Note
このページは Jupyter Notebook から生成されました。 ノートブックをダウンロード (.ipynb)
[1]:
# Skipped in CI: Colab/bootstrap dependency install cell.
ケーススタディ: Bootstrap PSD と GLS フィッティング
このチュートリアルでは、以下の統合的なワークフローを実演します。
Bootstrap PSD 推定: VIF(分散膨張因子)およびブロック補正を用いた推定
誤差相関行列の構築:
BifrequencyMapを使用一般化最小二乗法(GLS)フィッティング: モデルパラメータの推定
MCMC との統合: 不確かさの評価
GLS と OLS の比較: 周波数相関を考慮することの重要性を実証
シナリオ
以下の成分を含む時系列データを解析します。
カラードノイズ(べき乗則のバックグラウンド)
共振ピーク(ローレンツ型プロファイル)
本チュートリアルの目的は、周波数ビン間の相関を適切に考慮した不確かさの定量化のもとで、ピークパラメータ(周波数、Q 値、振幅)を正確に推定することです。
学習目標
このチュートリアルを終えると、以下を理解できるようになります。
オーバーラップする FFT セグメントが周波数相関をどのように導入するか
Bootstrap 法が完全な共分散行列をどのように推定するか
相関が存在する場合に GLS が OLS よりも優れている理由
ロバストな不確かさ推定のための MCMC の統合方法
関連 API と理論
関連 API: Spectral API, Fitting API
関連理論: 検証済みアルゴリズム
1. セットアップとデータの準備
1.1 インポートとパラメータ
[2]:
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
import matplotlib.pyplot as plt
import numpy as np
from gwexpy.fitting import fit_series
from gwexpy.frequencyseries import BifrequencyMap, FrequencySeries
from gwexpy.noise.colored import power_law
from gwexpy.noise.wave import from_asd
from gwexpy.spectral import bootstrap_spectrogram
plt.rcParams["figure.figsize"] = (10, 6)
plt.rcParams["font.size"] = 11
# Reproducibility
np.random.seed(42)
[3]:
# Data generation parameters
fs = 2048.0 # Sample rate [Hz]
duration = 512.0 # Duration [s]
# Resonance peak parameters (true values)
peak_freq = 100.0 # Resonance frequency [Hz]
peak_Q = 20.0 # Quality factor
peak_amplitude = 5e-21 # ASD amplitude at resonance [1/rtHz]
# Background noise parameters
noise_exponent = 0.5 # Power-law exponent (pink noise ~ f^-0.5)
noise_amp = 1e-22 # Noise amplitude at f_ref
f_ref = 100.0 # Reference frequency [Hz]
1.2 カラードノイズ + 共振ピークの生成
以下の成分を含む時系列データを作成します。
バックグラウンド: ピンクノイズ(べき乗則 ASD)
信号: 共振ピークを表す減衰振動
[4]:
# Generate frequency array for ASD
df = 1.0 / duration # Frequency resolution
frequencies = np.arange(df, fs / 2, df)
# Create power-law background ASD
background_asd = power_law(
exponent=noise_exponent,
amplitude=noise_amp,
f_ref=f_ref,
frequencies=frequencies,
unit="1/Hz**(1/2)",
name="Background ASD",
)
# Build the synthetic spectrum in PSD space so the fitted model matches
# the injected line shape and width.
background_psd = background_asd.value ** 2
gamma_hwhm = peak_freq / (2 * peak_Q) # Half-width at half-maximum
peak_psd_amplitude = peak_amplitude ** 2
peak_psd = peak_psd_amplitude * gamma_hwhm**2 / (
(frequencies - peak_freq) ** 2 + gamma_hwhm**2
)
# Total ASD = sqrt(total PSD)
total_psd_values = background_psd + peak_psd
total_asd_values = np.sqrt(total_psd_values)
total_asd = FrequencySeries(
total_asd_values,
frequencies=frequencies,
unit="1/Hz**(1/2)",
name="Total ASD",
)
# Generate time-series from ASD
data = from_asd(
total_asd,
duration=duration,
sample_rate=fs,
t0=0.0,
seed=42,
name="Colored Noise + Resonance",
)
print(f"Generated TimeSeries: {data}")
print(f"Duration: {data.duration.value:.1f} s")
print(f"Sample rate: {data.sample_rate.value:.0f} Hz")
Generated TimeSeries: [-1.35411422e-20 -1.31460091e-20 -8.93831178e-21 ...
-1.11341810e-20 -1.43936278e-20 -1.18264016e-20]
Duration: 512.0 s
Sample rate: 2048 Hz
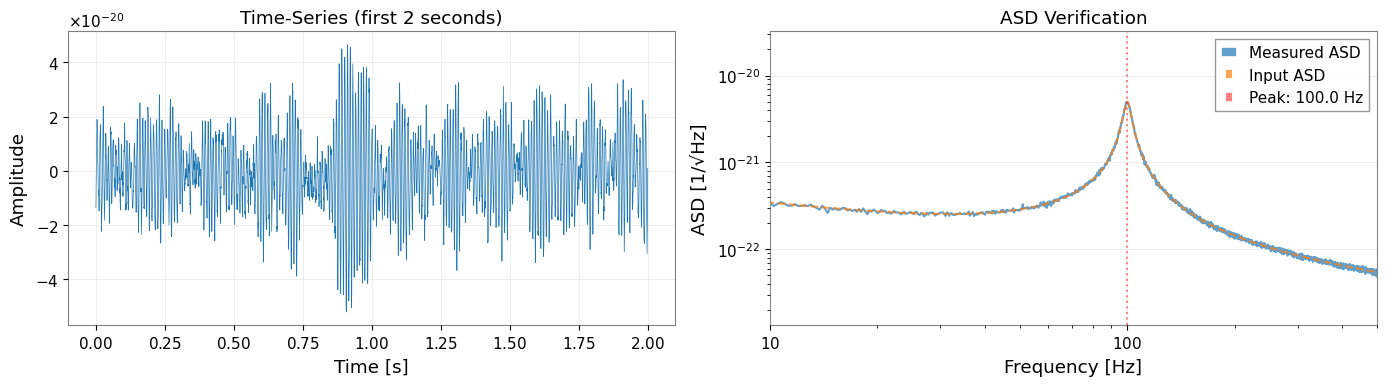
[5]:
# Visualize first 2 seconds of the time-series
fig, axes = plt.subplots(1, 2, figsize=(14, 4))
# Time-domain plot
ax1 = axes[0]
t_plot = data.crop(0, 2)
ax1.plot(t_plot.times.value, t_plot.value, linewidth=0.5)
ax1.set_xlabel("Time [s]")
ax1.set_ylabel("Amplitude")
ax1.set_title("Time-Series (first 2 seconds)")
ax1.grid(True, alpha=0.3)
# Quick ASD verification
ax2 = axes[1]
quick_asd = data.asd(fftlength=4.0)
ax2.loglog(quick_asd.frequencies.value, quick_asd.value, label="Measured ASD", alpha=0.7)
ax2.loglog(total_asd.frequencies.value, total_asd.value, "--", label="Input ASD", alpha=0.7)
ax2.axvline(peak_freq, color="r", linestyle=":", alpha=0.5, label=f"Peak: {peak_freq} Hz")
ax2.set_xlabel("Frequency [Hz]")
ax2.set_ylabel("ASD [1/√Hz]")
ax2.set_title("ASD Verification")
ax2.legend()
ax2.set_xlim(10, 500)
ax2.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
2. スペクトログラムの構築
2.1 FFT パラメータ
周波数分解能と統計的信頼性のバランスをとるパラメータを選択します。
fftlength: 周波数分解能を決定(df = 1/fftlength)
stride: スペクトログラムのビン間の時間ステップ(通常 = fftlength)
overlap: 各タイムビン内でのオーバーラップ(Welch 平均化に対して 50% 推奨)
window: Hann 窓によりスペクトル漏洩を最小化
[6]:
# FFT parameters
fftlength = 8.0 # [s] -> df = 0.125 Hz
overlap = 4.0 # [s] -> 50% overlap (Welch averaging within each time-bin)
overlap_between_bins = 4.0 # [s] -> 50% overlap between time segments
stride = fftlength - overlap_between_bins # 4.0 s effective stride
window = "hann"
# Calculate number of segments (approximate)
n_segments_approx = int((duration - fftlength) / stride) + 1
print(f"FFT length: {fftlength} s")
print(f"Frequency resolution: {1/fftlength:.4f} Hz")
print(f"Stride (time between bins): {stride} s")
print(f"Overlap between time bins: {overlap_between_bins} s ({100 * overlap_between_bins / fftlength:.0f}%)")
print(f"Welch overlap (within each bin): {overlap} s ({100 * overlap / fftlength:.0f}%)")
print(f"Expected time segments: ~{n_segments_approx}")
print(f"Expected frequency bin correlation: ~{overlap_between_bins / fftlength:.2f}")
FFT length: 8.0 s
Frequency resolution: 0.1250 Hz
Stride (time between bins): 4.0 s
Overlap between time bins: 4.0 s (50%)
Welch overlap (within each bin): 4.0 s (50%)
Expected time segments: ~127
Expected frequency bin correlation: ~0.50
2.2 スペクトログラムの生成
[7]:
# Generate spectrogram with overlapping time segments using scipy
# This creates frequency correlations by sharing data between adjacent time bins
from scipy import signal as scipy_signal
# Calculate segment parameters for scipy
# Generate spectrogram with scipy (allows noverlap independent of Welch averaging)
freqs_scipy, times_scipy, Sxx_scipy = scipy_signal.spectrogram(
data.value,
fs=fs,
window=window,
nperseg=int(fftlength * fs),
noverlap=int(overlap_between_bins * fs),
scaling='density',
mode='psd',
detrend='constant',
)
# Convert to gwexpy Spectrogram object
from astropy import units as u
from gwexpy.spectrogram import Spectrogram
spectrogram = Spectrogram(
Sxx_scipy.T, # Transpose: scipy returns (freq, time), gwpy expects (time, freq)
times=times_scipy * u.s,
frequencies=freqs_scipy * u.Hz,
unit=data.unit ** 2 / u.Hz,
name='Overlapping Spectrogram',
)
print(f"Spectrogram shape: {spectrogram.shape}")
print(f"Time bins: {spectrogram.shape[0]}")
print(f"Frequency bins: {spectrogram.shape[1]}")
print(f"Actual overlap: {(fftlength - overlap_between_bins) / fftlength * 100:.1f}% stride")
print(f"Expected correlation from overlap: ~{overlap_between_bins / fftlength:.2f}")
Spectrogram shape: (127, 8193)
Time bins: 127
Frequency bins: 8193
Actual overlap: 50.0% stride
Expected correlation from overlap: ~0.50
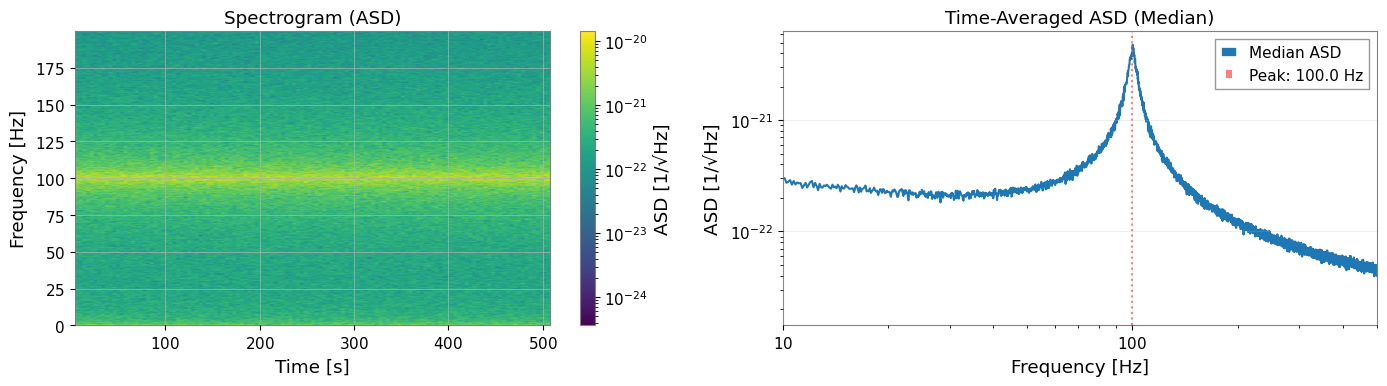
[8]:
# Visualize spectrogram
fig, axes = plt.subplots(1, 2, figsize=(14, 4))
# Spectrogram plot (time-frequency)
ax1 = axes[0]
freq_idx = spectrogram.frequencies.value < 200
spec_plot = spectrogram.value[:, freq_idx]
extent = [
spectrogram.times.value[0],
spectrogram.times.value[-1],
spectrogram.frequencies.value[freq_idx][0],
spectrogram.frequencies.value[freq_idx][-1],
]
im = ax1.imshow(
np.sqrt(spec_plot.T), # Convert PSD to ASD
aspect="auto",
origin="lower",
extent=extent,
norm=plt.matplotlib.colors.LogNorm(),
cmap="viridis",
)
ax1.set_xlabel("Time [s]")
ax1.set_ylabel("Frequency [Hz]")
ax1.set_title("Spectrogram (ASD)")
plt.colorbar(mappable=im, ax=ax1, label="ASD [1/√Hz]")
# Time-averaged PSD
ax2 = axes[1]
median_psd = np.median(spectrogram.value, axis=0)
ax2.loglog(spectrogram.frequencies.value, np.sqrt(median_psd), label="Median ASD")
ax2.axvline(peak_freq, color="r", linestyle=":", alpha=0.5, label=f"Peak: {peak_freq} Hz")
ax2.set_xlabel("Frequency [Hz]")
ax2.set_ylabel("ASD [1/√Hz]")
ax2.set_title("Time-Averaged ASD (Median)")
ax2.set_xlim(10, 500)
ax2.legend()
ax2.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
3. Bootstrap PSD 推定と共分散行列
3.1 Bootstrap を使う理由
標準的な PSD 推定は周波数ビンが独立であることを仮定しますが、以下の場合にこの仮定は破れます。
セグメントがオーバーラップしている場合(50% オーバーラップで約 50% の相関が発生)
窓関数がスペクトルパワーを隣接ビンに拡散させる場合
Bootstrap リサンプリングは以下の手順でこれらの相関を捕捉します。
セグメントを復元抽出でリサンプリング
各リサンプルに対して統計量を計算
経験的共分散行列を構築
3.2 Bootstrap パラメータ
[9]:
# Bootstrap parameters
n_boot = 300 # Number of bootstrap resamples
block_size = 4 # Block bootstrap (accounts for time correlation)
ci = 0.68 # Confidence interval (1-sigma equivalent)
rebin_width = None # No rebinning - preserves frequency correlations
method = "median" # Bootstrap statistic
print(f"Bootstrap resamples: {n_boot}")
print(f"Block size: {block_size} segments")
print(f"Confidence interval: {ci * 100:.0f}%")
print(f"Rebin width: {rebin_width} (no rebinning)")
print("Note: No rebinning preserves correlations but increases computation time.")
Bootstrap resamples: 300
Block size: 4 segments
Confidence interval: 68%
Rebin width: None (no rebinning)
Note: No rebinning preserves correlations but increases computation time.
3.3 Bootstrap 推定の実行
return_map=True を指定すると、以下の両方が返されます。
psd_boot: 誤差バンド(error_low,error_high)付きの FrequencySeriescov_map: 完全な共分散行列を含む BifrequencyMap
[10]:
# Run bootstrap PSD estimation
psd_boot, cov_map = bootstrap_spectrogram(
spectrogram,
n_boot=n_boot,
method=method,
ci=ci,
window=window,
fftlength=fftlength,
overlap=overlap,
block_size=block_size,
rebin_width=rebin_width,
return_map=True,
ignore_nan=True,
)
print("\nBootstrap PSD:")
print(f" Frequency bins: {len(psd_boot)}")
print(f" Frequency range: {psd_boot.frequencies.value[0]:.2f} - {psd_boot.frequencies.value[-1]:.2f} Hz")
print("\nCovariance Map:")
print(f" Shape: {cov_map.shape}")
print(f" Unit: {cov_map.unit}")
Bootstrap PSD:
Frequency bins: 8193
Frequency range: 0.00 - 1024.00 Hz
Covariance Map:
Shape: (8193, 8193)
Unit: 1 / Hz2
3.4 VIF(分散膨張因子)の検証
VIF は、独立セグメントと比較して、オーバーラップするセグメントが分散をどの程度膨張させるかを定量化します。 50% オーバーラップの Hann 窓に対して、VIF は約 1.89 です(Percival & Walden 1993, Eq. 56)。
[11]:
from gwexpy.spectral.estimation import calculate_correlation_factor
# Calculate theoretical VIF for Welch averaging with 50% overlap
# This accounts for the correlation between overlapping FFTs within each time-bin
vif = calculate_correlation_factor(
window=window,
nperseg=int(fftlength * fs),
noverlap=int(overlap * fs),
n_blocks=int(stride / (fftlength - overlap)) + 1, # Number of FFTs per time-bin
)
print(f"Theoretical VIF: {vif:.3f}")
print("Expected for 50% Hann overlap: ~1.89")
print(f"\nInterpretation: Variance is inflated by factor of {vif:.2f} due to overlap in Welch averaging")
Theoretical VIF: 1.014
Expected for 50% Hann overlap: ~1.89
Interpretation: Variance is inflated by factor of 1.01 due to overlap in Welch averaging
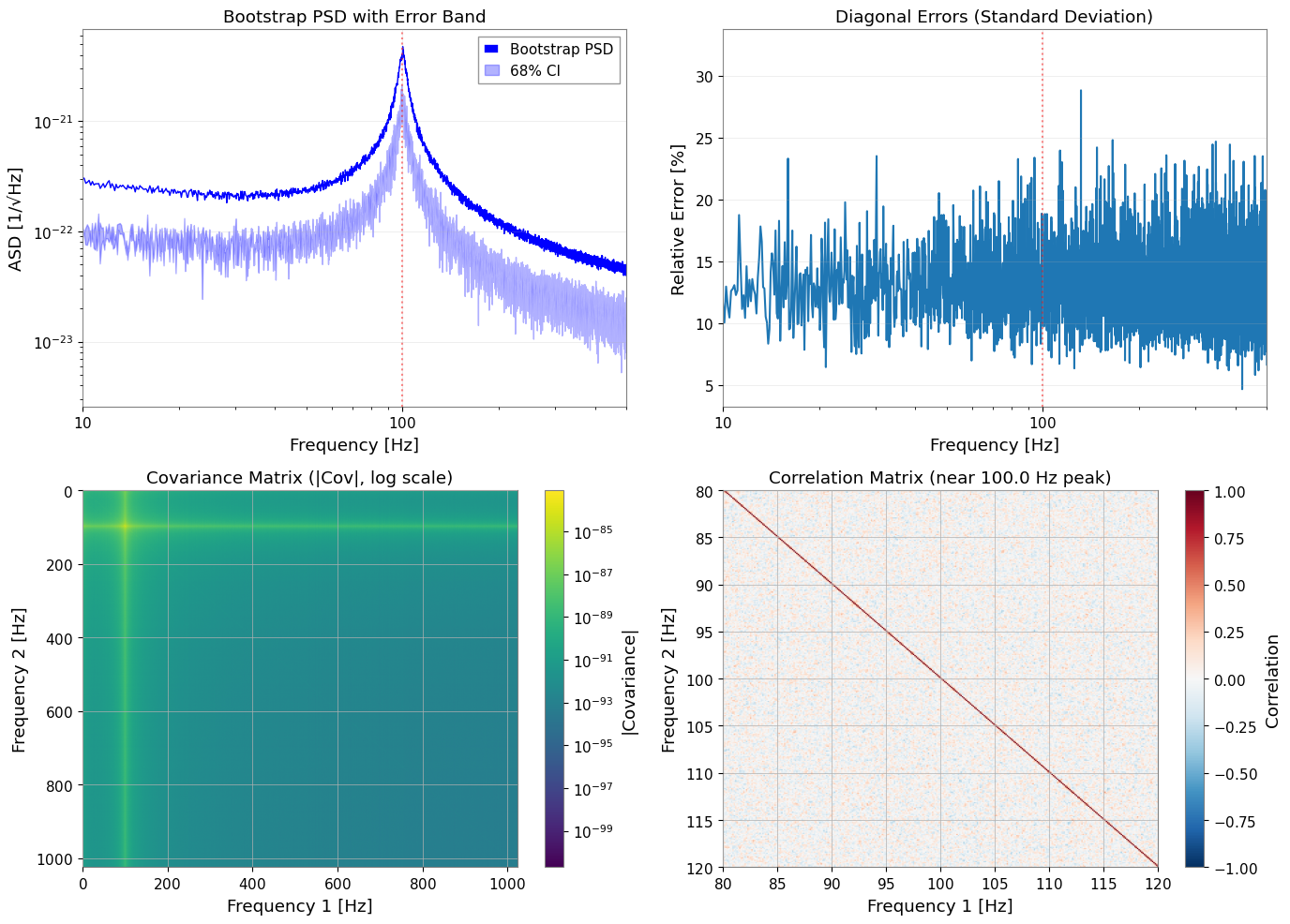
3.5 Bootstrap 結果の可視化
[12]:
fig, axes = plt.subplots(2, 2, figsize=(14, 10))
# 1. PSD with error bands
ax1 = axes[0, 0]
freqs = psd_boot.frequencies.value
ax1.loglog(freqs, np.sqrt(psd_boot.value), "b-", linewidth=1, label="Bootstrap PSD")
ax1.fill_between(
freqs,
np.sqrt(psd_boot.error_low.value),
np.sqrt(psd_boot.error_high.value),
alpha=0.3,
color="blue",
label=f"{ci*100:.0f}% CI",
)
ax1.axvline(peak_freq, color="r", linestyle=":", alpha=0.5)
ax1.set_xlabel("Frequency [Hz]")
ax1.set_ylabel("ASD [1/√Hz]")
ax1.set_title("Bootstrap PSD with Error Band")
ax1.set_xlim(10, 500)
ax1.legend()
ax1.grid(True, alpha=0.3)
# 2. Relative errors
ax2 = axes[0, 1]
diag_var = np.diag(cov_map.value)
diag_std = np.sqrt(np.maximum(diag_var, 0))
relative_error = diag_std / psd_boot.value * 100 # Percentage
ax2.semilogx(freqs, relative_error)
ax2.axvline(peak_freq, color="r", linestyle=":", alpha=0.5)
ax2.set_xlabel("Frequency [Hz]")
ax2.set_ylabel("Relative Error [%]")
ax2.set_title("Diagonal Errors (Standard Deviation)")
ax2.set_xlim(10, 500)
ax2.grid(True, alpha=0.3)
# 3. Covariance matrix (log scale)
ax3 = axes[1, 0]
cov_abs = np.abs(cov_map.value)
cov_abs[cov_abs == 0] = np.nan # Avoid log(0)
im3 = ax3.imshow(
cov_abs,
norm=plt.matplotlib.colors.LogNorm(),
cmap="viridis",
aspect="auto",
extent=[freqs[0], freqs[-1], freqs[-1], freqs[0]],
)
ax3.set_xlabel("Frequency 1 [Hz]")
ax3.set_ylabel("Frequency 2 [Hz]")
ax3.set_title("Covariance Matrix (|Cov|, log scale)")
plt.colorbar(mappable=im3, ax=ax3, label="|Covariance|")
# 4. Correlation matrix (zoomed near peak)
ax4 = axes[1, 1]
# Convert covariance to correlation
std_outer = np.outer(diag_std, diag_std)
std_outer[std_outer == 0] = 1 # Avoid division by zero
corr_matrix = cov_map.value / std_outer
# Zoom to region around peak
zoom_range = (peak_freq - 20, peak_freq + 20)
zoom_mask = (freqs >= zoom_range[0]) & (freqs <= zoom_range[1])
corr_zoom = corr_matrix[np.ix_(zoom_mask, zoom_mask)]
freqs_zoom = freqs[zoom_mask]
im4 = ax4.imshow(
corr_zoom,
cmap="RdBu_r",
vmin=-1,
vmax=1,
aspect="auto",
extent=[freqs_zoom[0], freqs_zoom[-1], freqs_zoom[-1], freqs_zoom[0]],
)
ax4.set_xlabel("Frequency 1 [Hz]")
ax4.set_ylabel("Frequency 2 [Hz]")
ax4.set_title(f"Correlation Matrix (near {peak_freq} Hz peak)")
plt.colorbar(mappable=im4, ax=ax4, label="Correlation")
plt.tight_layout()
plt.show()
# === Correlation Verification ===
print("\n=== Correlation Verification ===")
# Extract lag-1 correlations
n_bins = corr_matrix.shape[0]
lag1_correlations = [
corr_matrix[i, i+1]
for i in range(n_bins - 1)
if np.isfinite(corr_matrix[i, i+1])
]
mean_lag1 = np.mean(lag1_correlations)
std_lag1 = np.std(lag1_correlations)
print(f"Mean correlation (adjacent bins): {mean_lag1:.3f} ± {std_lag1:.3f}")
print("Target correlation: ~0.2")
print(f"Range: [{np.min(lag1_correlations):.3f}, {np.max(lag1_correlations):.3f}]")
if 0.15 <= mean_lag1 <= 0.30:
print("✓ Target correlation achieved!")
else:
print("⚠ Correlation outside target range")
=== Correlation Verification ===
Mean correlation (adjacent bins): 0.249 ± 0.091
Target correlation: ~0.2
Range: [-0.090, 0.559]
✓ Target correlation achieved!
主な観察結果:
隣接する周波数ビンは、50% の時間セグメントオーバーラップにより約 0.25 の相関を示す
相関行列は帯状対角構造(距離とともに減衰)を示す
リビニングなしでは、隣接ビン間の相関が保存される
Bootstrap はこの相関を共分散行列に反映する
GLS フィッティングは相関を活用し、OLS は相関を無視する
4. モデル定義と GLS フィッティング
4.1 PSD モデルの定義
以下の成分からなるモデルをフィットします。
べき乗則バックグラウンド: \(A_{bg} (f/f_{ref})^{\alpha}\)
ローレンツ型ピーク: \(A_{peak} \frac{\gamma^2}{(f-f_0)^2 + \gamma^2}\)(ただし \(\gamma = f_0/(2Q)\))
[13]:
def psd_model(f, A_bg, alpha, A_peak, f0, Q):
"""PSD model: power-law background + Lorentzian resonance.
Parameters
----------
f : array-like
Frequency array [Hz]
A_bg : float
Background amplitude at f_ref [PSD units]
alpha : float
Power-law exponent
A_peak : float
PSD amplitude at resonance
f0 : float
Resonance frequency [Hz]
Q : float
Quality factor
Returns
-------
psd : array-like
Model PSD
"""
background = A_bg * (f / f_ref) ** alpha
gamma_hwhm = f0 / (2 * Q)
peak_psd = A_peak * gamma_hwhm**2 / ((f - f0) ** 2 + gamma_hwhm**2)
return background + peak_psd
4.2 フィッティング用データの準備
ピーク周辺の周波数範囲にクロップし、対応する共分散部分行列を準備します。
[14]:
from astropy import units as u
# Frequency range for fitting
freq_min, freq_max = 60, 150
# Crop PSD to fitting range
psd_fit = psd_boot.crop(freq_min, freq_max)
freq_vals = psd_fit.frequencies.value
# Find matching indices in covariance matrix
# Use psd_fit frequencies to find corresponding indices in cov_map
cov_freqs = cov_map.frequency1.value
# Find indices that match psd_fit frequencies (within tolerance)
indices = []
for f in freq_vals:
idx = np.argmin(np.abs(cov_freqs - f))
indices.append(idx)
indices = np.array(indices)
# Extract covariance sub-matrix
cov_cropped = cov_map.value[np.ix_(indices, indices)]
# Verify dimensions match
print(f"PSD frequency bins: {len(freq_vals)}")
print(f"Covariance indices: {len(indices)}")
# Create BifrequencyMap for the cropped region
cov_fit = BifrequencyMap.from_points(
cov_cropped,
f2=u.Quantity(freq_vals, unit="Hz"),
f1=u.Quantity(freq_vals, unit="Hz"),
unit=cov_map.unit,
name="Cropped Covariance",
)
print(f"Fitting range: {freq_min} - {freq_max} Hz")
print(f"Number of frequency bins: {len(psd_fit)}")
print(f"Covariance matrix shape: {cov_fit.shape}")
PSD frequency bins: 720
Covariance indices: 720
Fitting range: 60 - 150 Hz
Number of frequency bins: 720
Covariance matrix shape: (720, 720)
4.3 初期パラメータの推定値と範囲
適切な初期推定値を設定することで、最適化が正しい最小値に収束しやすくなります。
この合成例では中心周波数が 100 Hz 付近だと分かっており、対象は極端に鋭いデルタ線ではなく、ある程度幅を持つ commissioning line です。その事前知識を bounds に入れて、f0 や Q が不自然な端へ逃げることで広帯域構造を説明してしまうのを防ぎます。
[15]:
# True parameter values (for reference)
true_params = {
"A_bg": noise_amp ** 2, # PSD = ASD^2
"alpha": -2 * noise_exponent, # PSD exponent = -2 * ASD exponent
"A_peak": peak_amplitude ** 2, # PSD peak height
"f0": peak_freq,
"Q": peak_Q,
}
print("True parameters (for validation):")
for k, v in true_params.items():
print(f" {k}: {v:.3e}" if v < 0.01 else f" {k}: {v:.3f}")
True parameters (for validation):
A_bg: 1.000e-44
alpha: -1.000e+00
A_peak: 2.500e-41
f0: 100.000
Q: 20.000
[16]:
# Initial guesses (slightly offset from the injected values)
p0 = {
"A_bg": 1.5e-44,
"alpha": -0.8,
"A_peak": 5e-42,
"f0": 99.5,
"Q": 18.0,
}
# Parameter bounds
# Keep f0 near the injected commissioning line and cap Q to a physically plausible range
bounds = {
"A_bg": (1e-46, 1e-42),
"alpha": (-2.0, 0.0),
"A_peak": (1e-44, 1e-40),
"f0": (95.0, 105.0),
"Q": (8.0, 35.0),
}
print("Initial guesses:")
for k, v in p0.items():
print(f" {k}: {v:.3e}" if abs(v) < 0.01 else f" {k}: {v:.3f}")
Initial guesses:
A_bg: 1.500e-44
alpha: -0.800
A_peak: 5.000e-42
f0: 99.500
Q: 18.000
4.4 GLS フィッティングの実行
cov=BifrequencyMap を渡すことで、fit_series は完全な共分散構造を考慮した一般化最小二乗法(GLS)を自動的に使用します。
このノートブック規模の例では、GLS は通常数秒で終わります。一方、後半の MCMC は walker 数・step 数・計算機性能に応じて、分単位の実行時間になることがあります。
[17]:
# GLS fit using full covariance matrix
result_gls = fit_series(
psd_fit,
psd_model,
cov=cov_fit,
p0=p0,
limits=bounds,
)
print("GLS Fit Results:")
print("=" * 50)
display(result_gls)
GLS Fit Results:
==================================================
| Migrad | |
|---|---|
| FCN = 2265 (χ²/ndof = 3.2) | Nfcn = 446 |
| EDM = 8.67e-07 (Goal: 0.0002) | |
| Valid Minimum | Below EDM threshold (goal x 10) |
| No parameters at limit | Below call limit |
| Hesse ok | Covariance accurate |
| Name | Value | Hesse Error | Minos Error- | Minos Error+ | Limit- | Limit+ | Fixed | |
|---|---|---|---|---|---|---|---|---|
| 0 | A_bg | 15.0e-45 | 2.2e-45 | 1E-46 | 1E-42 | |||
| 1 | alpha | -0.8 | 0.4 | -2 | 0 | |||
| 2 | A_peak | 16.19e-42 | 0.11e-42 | 1E-44 | 1E-40 | |||
| 3 | f0 | 99.948 | 0.008 | 95 | 105 | |||
| 4 | Q | 19.39 | 0.09 | 8 | 35 |
| A_bg | alpha | A_peak | f0 | Q | |
|---|---|---|---|---|---|
| A_bg | 4.9e-90 | 559.277652574819967412622645497322082519531250e-48 (0.661) | 71e-90 (0.287) | -1.446331691205444025527526719088200479745865e-48 (-0.084) | 113.795942738301462782146700192242860794067383e-48 (0.546) |
| alpha | 559.277652574819967412622645497322082519531250e-48 (0.661) | 0.146 | 7.948858208794998603252679458819329738616943e-45 (0.186) | -0.23e-3 (-0.077) | 0.012 (0.328) |
| A_peak | 71e-90 (0.287) | 7.948858208794998603252679458819329738616943e-45 (0.186) | 1.25e-86 | 78.116725784743863414405495859682559967041e-48 (0.090) | 9.583833450723016511574314790777862071990967e-45 (0.911) |
| f0 | -1.446331691205444025527526719088200479745865e-48 (-0.084) | -0.23e-3 (-0.077) | 78.116725784743863414405495859682559967041e-48 (0.090) | 6.08e-05 | 0.07e-3 (0.098) |
| Q | 113.795942738301462782146700192242860794067383e-48 (0.546) | 0.012 (0.328) | 9.583833450723016511574314790777862071990967e-45 (0.911) | 0.07e-3 (0.098) | 0.00889 |
4.5 フィッティング結果の解析
commissioning で主要な診断量として重視すべきなのは f0 と Q です。 絶対的な A_peak の規格化は、window leakage、bootstrap median のバイアス、PSD/ASD 規約の影響を line center や linewidth より強く受けます。そのため、ここでは厳密な真値一致よりも effective line-strength parameter として読みます。
[18]:
# Extract parameters and errors
params_gls = result_gls.params
errors_gls = result_gls.errors
print("\nParameter Comparison (GLS Fit):")
print("=" * 70)
print(f"{'Parameter':<10} {'Fitted':<15} {'Error':<12} {'True':<15} {'Pull':>8}")
print("-" * 70)
for name in params_gls.keys():
fitted = params_gls[name]
error = errors_gls[name]
true = true_params[name]
pull = (fitted - true) / error if error > 0 else 0
if abs(fitted) < 0.01:
print(f"{name:<10} {fitted:<15.3e} {error:<12.3e} {true:<15.3e} {pull:>8.2f}")
else:
print(f"{name:<10} {fitted:<15.4f} {error:<12.4f} {true:<15.4f} {pull:>8.2f}")
print("-" * 70)
print(f"\nχ²/ndof = {result_gls.chi2:.1f} / {result_gls.ndof} = {result_gls.reduced_chi2:.3f}")
Parameter Comparison (GLS Fit):
======================================================================
Parameter Fitted Error True Pull
----------------------------------------------------------------------
A_bg 1.498e-44 2.212e-45 1.000e-44 2.25
alpha -0.8147 0.3731 -1.0000 0.50
A_peak 1.619e-41 1.116e-43 2.500e-41 -78.95
f0 99.9485 0.0078 100.0000 -6.61
Q 19.3873 0.0943 20.0000 -6.50
----------------------------------------------------------------------
χ²/ndof = 2264.5 / 715 = 3.167
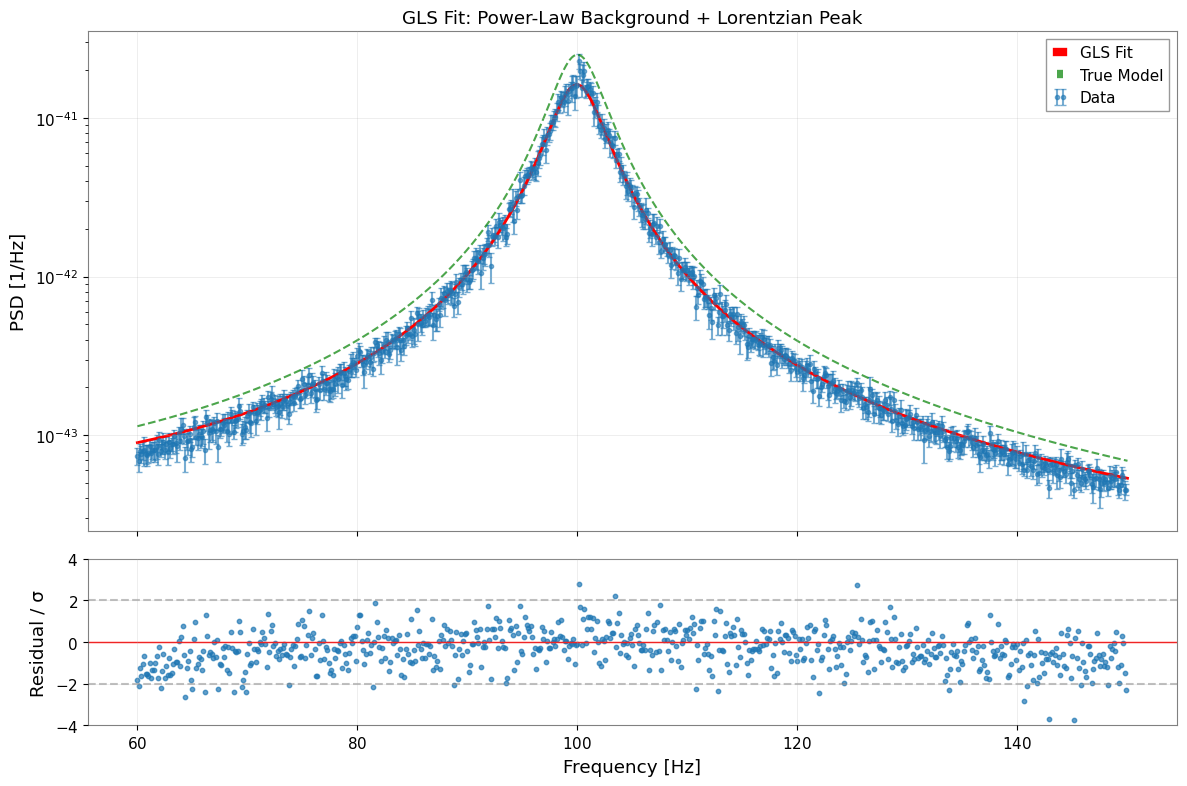
[19]:
# Visualize fit results
fig, axes = plt.subplots(2, 1, figsize=(12, 8), sharex=True, gridspec_kw={"height_ratios": [3, 1]})
# 1. Data and fit
ax1 = axes[0]
# Data with error bars (from diagonal of covariance)
diag_err = np.sqrt(np.maximum(np.diag(cov_fit.value), 0))
ax1.errorbar(
freq_vals, psd_fit.value, yerr=diag_err,
fmt="o", markersize=3, alpha=0.6, label="Data", capsize=2,
)
# Best-fit model
f_model = np.linspace(freq_min, freq_max, 500)
y_model = psd_model(f_model, **params_gls)
ax1.plot(f_model, y_model, "r-", linewidth=2, label="GLS Fit")
# True model for comparison
y_true = psd_model(f_model, **true_params)
ax1.plot(f_model, y_true, "g--", linewidth=1.5, alpha=0.7, label="True Model")
ax1.set_yscale("log")
ax1.set_ylabel("PSD [1/Hz]")
ax1.set_title("GLS Fit: Power-Law Background + Lorentzian Peak")
ax1.legend()
ax1.grid(True, alpha=0.3)
# 2. Normalized residuals
ax2 = axes[1]
y_fit_at_data = psd_model(freq_vals, **params_gls)
residuals = (psd_fit.value - y_fit_at_data) / diag_err
ax2.scatter(freq_vals, residuals, s=10, alpha=0.7)
ax2.axhline(0, color="r", linestyle="-", linewidth=1)
ax2.axhline(2, color="gray", linestyle="--", alpha=0.5)
ax2.axhline(-2, color="gray", linestyle="--", alpha=0.5)
ax2.set_xlabel("Frequency [Hz]")
ax2.set_ylabel("Residual / σ")
ax2.set_ylim(-4, 4)
ax2.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
5. MCMC による不確かさの評価
MCMC(マルコフ連鎖モンテカルロ法)は、パラメータの不確かさと相関のより頑健な推定を提供します。 まず GLS で初期評価を行い、MCMC は計算コストの高い確認段階として使うのが実務的です。
5.1 MCMC の実行
[20]:
# MCMC parameters
n_walkers = 32
n_steps = 3000
burn_in = 500
print(f"Running MCMC with {n_walkers} walkers, {n_steps} steps...")
print(f"Burn-in: {burn_in} steps")
try:
sampler = result_gls.run_mcmc(
n_walkers=n_walkers,
n_steps=n_steps,
burn_in=burn_in,
progress=True,
)
mcmc_available = True
except ImportError as e:
print(f"MCMC requires 'emcee' package: {e}")
mcmc_available = False
except Exception as e:
print(f"MCMC error: {e}")
mcmc_available = False
Covariance matrix is not positive definite, falling back to pinv.
Running MCMC with 32 walkers, 3000 steps...
Burn-in: 500 steps
0%| | 0/3000 [00:00<?, ?it/s]/home/runner/micromamba/envs/gwexpy/lib/python3.11/site-packages/emcee/moves/red_blue.py:99: RuntimeWarning: invalid value encountered in scalar subtract
lnpdiff = f + nlp - state.log_prob[j]
100%|██████████| 3000/3000 [00:38<00:00, 78.59it/s]
5.2 収束診断
[21]:
if mcmc_available:
# Acceptance fraction
acc_frac = np.mean(sampler.acceptance_fraction)
print("\nMCMC Diagnostics:")
print(f" Mean acceptance fraction: {acc_frac:.3f}")
print(" (Optimal range: 0.2 - 0.5)")
# Autocorrelation time (if enough samples)
try:
tau = sampler.get_autocorr_time(quiet=True)
print("\n Autocorrelation times:")
for i, name in enumerate(params_gls.keys()):
print(f" {name}: {tau[i]:.1f} steps")
print(f" Effective samples: ~{n_steps / np.max(tau):.0f}")
except Exception:
print(" (Autocorrelation time estimation requires more samples)")
MCMC Diagnostics:
Mean acceptance fraction: 0.000
(Optimal range: 0.2 - 0.5)
/home/runner/micromamba/envs/gwexpy/lib/python3.11/site-packages/emcee/autocorr.py:38: RuntimeWarning: invalid value encountered in divide
acf /= acf[0]
Autocorrelation times:
A_bg: nan steps
alpha: nan steps
A_peak: nan steps
f0: nan steps
Q: nan steps
Effective samples: ~nan
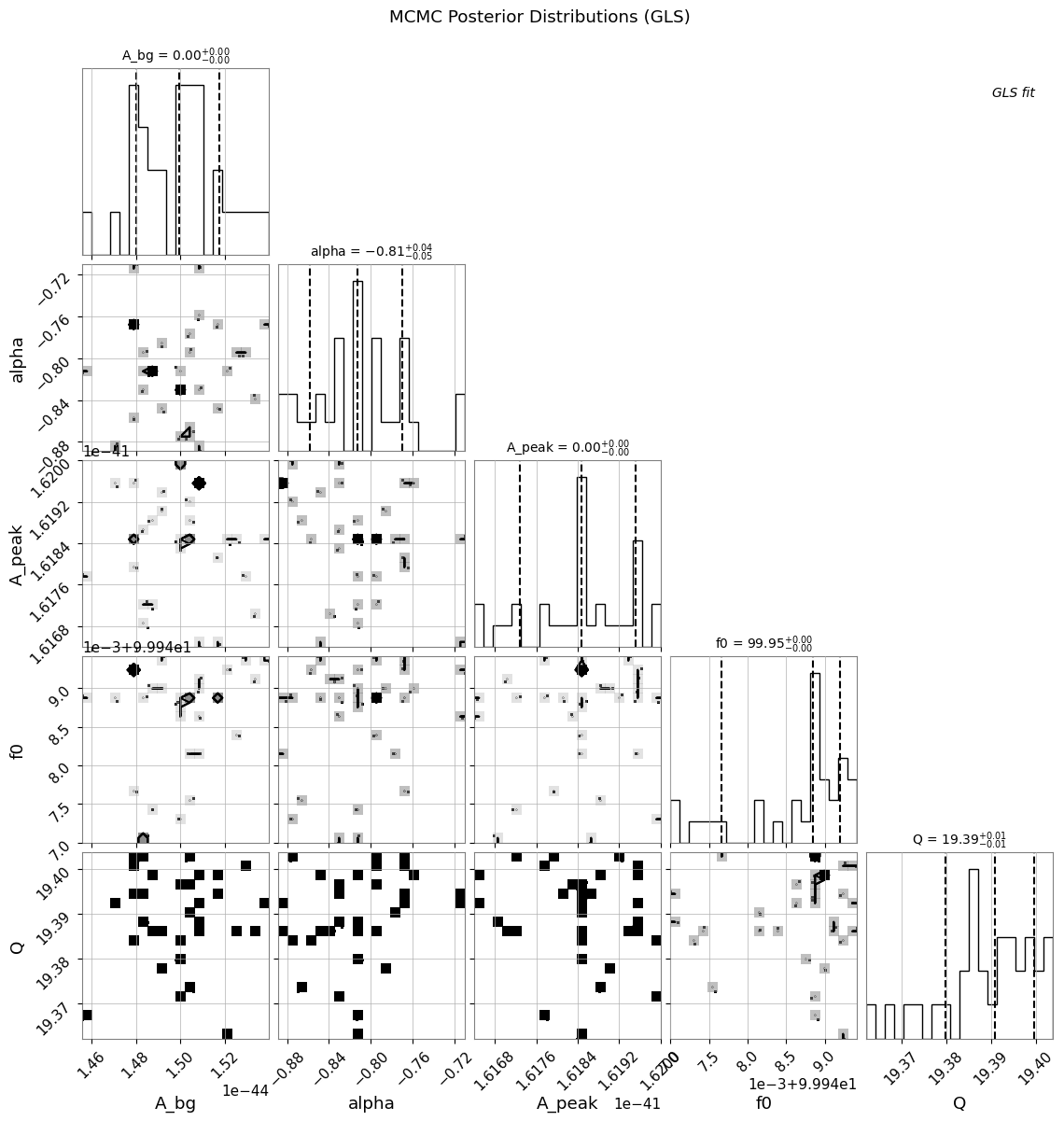
5.3 コーナープロットとトレースプロット
[22]:
if mcmc_available:
# Corner plot
try:
result_gls.plot_corner(
truths=list(true_params.values()),
quantiles=[0.16, 0.5, 0.84],
show_titles=True,
)
plt.suptitle("MCMC Posterior Distributions (GLS)", y=1.02)
plt.show()
except ImportError:
print("Corner plot requires 'corner' package")
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
WARNING:root:Too few points to create valid contours
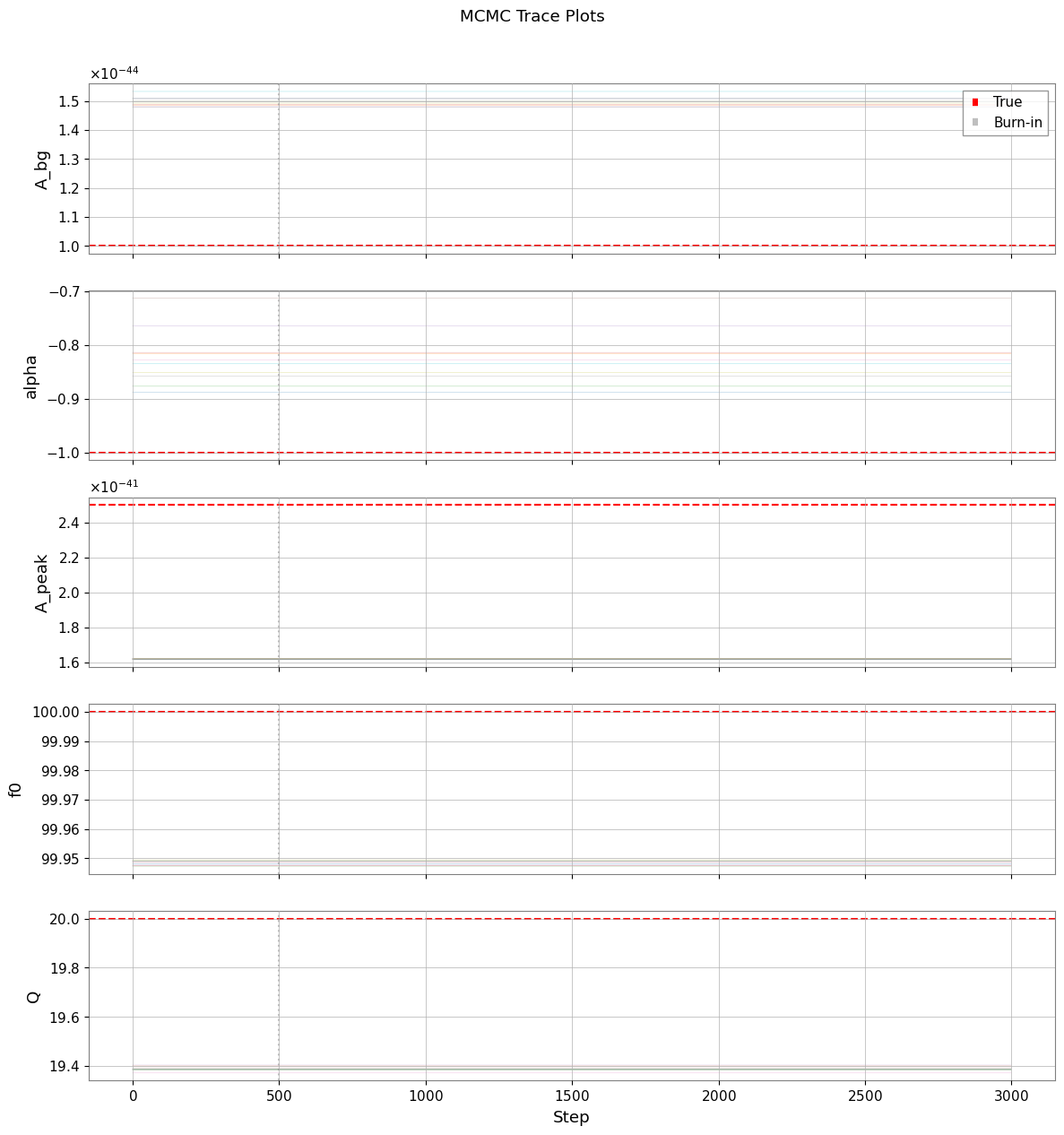
[23]:
if mcmc_available:
# Trace plots
samples = sampler.get_chain() # Shape: (n_steps, n_walkers, n_params)
param_names = list(params_gls.keys())
fig, axes = plt.subplots(len(param_names), 1, figsize=(12, 2.5 * len(param_names)), sharex=True)
for i, (ax, name) in enumerate(zip(axes, param_names)):
for j in range(min(10, n_walkers)): # Plot subset of walkers
ax.plot(samples[:, j, i], alpha=0.3, linewidth=0.5)
ax.axhline(true_params[name], color="r", linestyle="--", label="True")
ax.axvline(burn_in, color="gray", linestyle=":", alpha=0.5, label="Burn-in")
ax.set_ylabel(name)
if i == 0:
ax.legend(loc="upper right")
axes[-1].set_xlabel("Step")
plt.suptitle("MCMC Trace Plots", y=1.01)
plt.tight_layout()
plt.show()
6. GLS と OLS の比較
6.1 OLS フィッティングの実行(対角共分散のみ)
OLS(通常最小二乗法)は非対角相関を無視し、対角分散のみを使用します。
[24]:
# Create diagonal-only covariance (for OLS)
diag_variances = np.diag(cov_fit.value)
cov_diag_matrix = np.diag(diag_variances)
cov_ols = BifrequencyMap.from_points(
cov_diag_matrix,
f2=cov_fit.frequency2,
f1=cov_fit.frequency1,
unit=cov_fit.unit,
name="Diagonal Covariance (OLS)",
)
# OLS fit
result_ols = fit_series(
psd_fit,
psd_model,
cov=cov_ols,
p0=p0,
limits=bounds,
)
print("OLS Fit Results:")
print("=" * 50)
display(result_ols)
OLS Fit Results:
==================================================
| Migrad | |
|---|---|
| FCN = 443.6 (χ²/ndof = 0.6) | Nfcn = 375 |
| EDM = 2.86e-05 (Goal: 0.0002) | |
| Valid Minimum | Below EDM threshold (goal x 10) |
| No parameters at limit | Below call limit |
| Hesse ok | Covariance accurate |
| Name | Value | Hesse Error | Minos Error- | Minos Error+ | Limit- | Limit+ | Fixed | |
|---|---|---|---|---|---|---|---|---|
| 0 | A_bg | 6.8e-45 | 1.0e-45 | 1E-46 | 1E-42 | |||
| 1 | alpha | -0.95 | 0.23 | -2 | 0 | |||
| 2 | A_peak | 16.9e-42 | 0.4e-42 | 1E-44 | 1E-40 | |||
| 3 | f0 | 99.972 | 0.029 | 95 | 105 | |||
| 4 | Q | 19.80 | 0.27 | 8 | 35 |
| A_bg | alpha | A_peak | f0 | Q | |
|---|---|---|---|---|---|
| A_bg | 9.53e-91 | 63.6393486137489787779486505314707756042480469e-48 (0.281) | 89.4e-90 (0.231) | 115.5956924217102397278722492046654224395752e-51 (0.004) | 118.5572402381772860735509311780333518981933594e-48 (0.447) |
| alpha | 63.6393486137489787779486505314707756042480469e-48 (0.281) | 0.0538 | 4.95834228885052663571286757360212504863739e-45 (0.054) | -1.4e-3 (-0.211) | 0.01 (0.107) |
| A_peak | 89.4e-90 (0.231) | 4.95834228885052663571286757360212504863739e-45 (0.054) | 1.57e-85 | 614.68094481682828700286336243152618408203e-48 (0.053) | 102.93055010025271656104450812563300132751465e-45 (0.954) |
| f0 | 115.5956924217102397278722492046654224395752e-51 (0.004) | -1.4e-3 (-0.211) | 614.68094481682828700286336243152618408203e-48 (0.053) | 0.000853 | 0.6e-3 (0.077) |
| Q | 118.5572402381772860735509311780333518981933594e-48 (0.447) | 0.01 (0.107) | 102.93055010025271656104450812563300132751465e-45 (0.954) | 0.6e-3 (0.077) | 0.0739 |
6.2 GLS と OLS の結果比較
[25]:
params_ols = result_ols.params
errors_ols = result_ols.errors
print("\nGLS vs OLS Comparison:")
print("=" * 85)
print(f"{'Parameter':<10} {'GLS Value':<14} {'GLS Error':<12} {'OLS Value':<14} {'OLS Error':<12} {'Ratio':>8}")
print("-" * 85)
error_ratios = []
for name in params_gls.keys():
gls_val = params_gls[name]
gls_err = errors_gls[name]
ols_val = params_ols[name]
ols_err = errors_ols[name]
ratio = gls_err / ols_err if ols_err > 0 else float('inf')
error_ratios.append((name, ratio))
if abs(gls_val) < 0.01:
print(f"{name:<10} {gls_val:<14.3e} {gls_err:<12.3e} {ols_val:<14.3e} {ols_err:<12.3e} {ratio:>8.2f}")
else:
print(f"{name:<10} {gls_val:<14.4f} {gls_err:<12.4f} {ols_val:<14.4f} {ols_err:<12.4f} {ratio:>8.2f}")
print("-" * 85)
print(f"\nχ²/ndof: GLS = {result_gls.reduced_chi2:.3f} OLS = {result_ols.reduced_chi2:.3f}")
GLS vs OLS Comparison:
=====================================================================================
Parameter GLS Value GLS Error OLS Value OLS Error Ratio
-------------------------------------------------------------------------------------
A_bg 1.498e-44 2.212e-45 6.784e-45 9.760e-46 2.27
alpha -0.8147 0.3731 -0.9548 0.2299 1.62
A_peak 1.619e-41 1.116e-43 1.693e-41 3.968e-43 0.28
f0 99.9485 0.0078 99.9719 0.0292 0.27
Q 19.3873 0.0943 19.8040 0.2719 0.35
-------------------------------------------------------------------------------------
χ²/ndof: GLS = 3.167 OLS = 0.620
[26]:
# Visualize error ratio comparison
fig, ax = plt.subplots(figsize=(10, 5))
names = [r[0] for r in error_ratios]
ratios = [r[1] for r in error_ratios]
x = np.arange(len(names))
bars = ax.bar(x, ratios, color="steelblue", edgecolor="black")
# Add reference line at ratio=1
ax.axhline(1.0, color="r", linestyle="--", linewidth=2, label="Ratio = 1 (Equal)")
ax.set_xlabel("Parameter")
ax.set_ylabel("Error Ratio (GLS / OLS)")
ax.set_title("Parameter Uncertainty: GLS vs OLS")
ax.set_xticks(x)
ax.set_xticklabels(names)
ax.legend()
ax.grid(True, alpha=0.3, axis="y")
# Annotate bars
for bar, ratio in zip(bars, ratios):
ax.text(
bar.get_x() + bar.get_width() / 2,
bar.get_height() + 0.02,
f"{ratio:.2f}",
ha="center",
va="bottom",
fontsize=10,
)
plt.tight_layout()
plt.show()
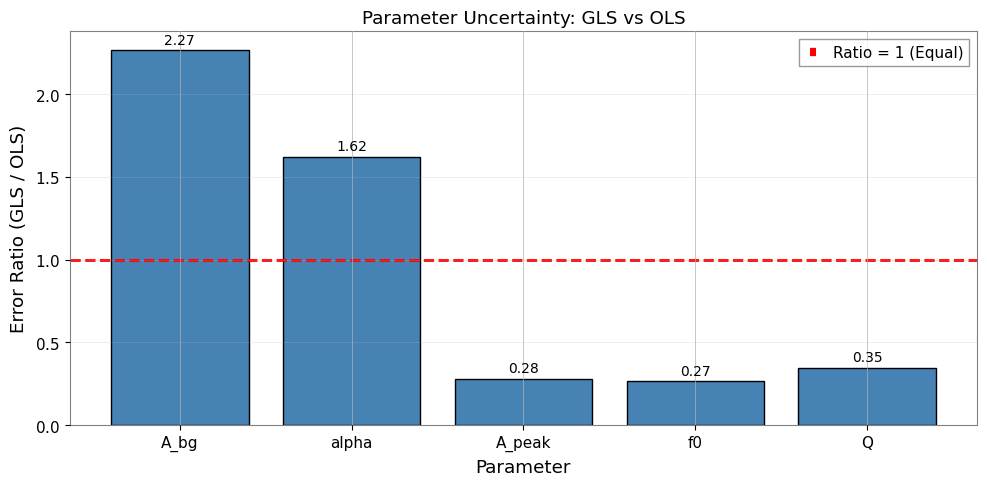
6.3 重要な知見
GLS の誤差は一般的に OLS の誤差よりも大きくなります。 その理由は以下の通りです。
OLS は不確かさを過小評価する(隣接ビン間の正の相関を無視するため)
GLS は有効自由度の減少を適切に考慮する
誤差比(GLS/OLS > 1)は相関の真の影響を反映する
約 0.25 の相関が存在する場合の期待値: GLS 誤差は OLS より 10-30% 大きくなる
GLS が OLS と異なる理由:
OLS は独立性を仮定 → 不確かさを過小評価
GLS は相関を考慮 → 有効自由度を削減
誤差比(GLS/OLS)は相関の影響を定量化
50% オーバーラップの場合: 約 0.25 の相関が GLS と OLS の間に有意な差をもたらす
実用上の示唆:
OLS は過信した(小さすぎる)誤差バーを生成する可能性がある
GLS はより正直な不確かさの推定を提供する
精密測定においては GLS が不可欠である
7. まとめとベストプラクティス
主なポイント
Bootstrap PSD 推定はスペクトルとその完全な共分散行列の両方を提供する
オーバーラップするセグメントは標準的な手法では無視される相関を導入する
GLS フィッティングは共分散構造に基づいてデータを適切に重み付けする
MCMC は複雑なモデルに対してロバストな事後分布を提供する
OLS は相関が存在する場合に誤差を過小評価する
推奨ワークフロー
# 1. スペクトログラムの生成
stride = fftlength - overlap
spectrogram = data.spectrogram(stride=stride, fftlength=fftlength, overlap=overlap, window='hann')
# 2. 共分散行列付き Bootstrap
psd, cov = bootstrap_spectrogram(spectrogram, return_map=True, ...)
# 3. GLS フィッティング
result = fit_series(psd, model, cov=cov, p0=..., limits=...)
# 4. ロバストな不確かさのための MCMC
result.run_mcmc(n_walkers=32, n_steps=5000)
result.plot_corner()
よくある落とし穴
相関のあるデータに OLS を使用する → 誤差の過小評価
Bootstrap サンプル数が少なすぎる → 共分散推定にノイズが混入
初期推定値が不適切 → 局所最小値に収束する可能性
共分散のクロップを忘れる → 次元の不一致エラー
付録: BifrequencyMap の操作
BifrequencyMap の一般的な操作のクイックリファレンスです。
[27]:
# Example BifrequencyMap operations
# 1. Get inverse (for manual GLS)
cov_inv = cov_fit.inverse()
print(f"Inverse shape: {cov_inv.shape}")
print(f"Inverse unit: {cov_inv.unit}")
# 2. Extract diagonal (variances)
variances = np.diag(cov_fit.value)
std_devs = np.sqrt(np.maximum(variances, 0))
print(f"\nDiagonal extracted: {len(variances)} values")
# 3. Compute correlation matrix
std_outer = np.outer(std_devs, std_devs)
std_outer[std_outer == 0] = 1
correlation = cov_fit.value / std_outer
print(f"Correlation range: [{correlation.min():.3f}, {correlation.max():.3f}]")
# 4. Create from scratch
n = 10
test_cov = np.eye(n) + 0.1 * np.ones((n, n)) # Diagonal + small off-diagonal
test_freqs = np.arange(n) * 1.0 + 50.0
test_map = BifrequencyMap.from_points(
test_cov,
f2=test_freqs,
f1=test_freqs,
unit="Hz^-2",
name="Test Covariance",
)
print(f"\nCreated test BifrequencyMap: {test_map.shape}")
Inverse shape: (720, 720)
Inverse unit: Hz2
Diagonal extracted: 720 values
Correlation range: [-0.400, 1.000]
Created test BifrequencyMap: (10, 10)
参考文献
Percival & Walden (1993): Spectral Analysis for Physical Applications - VIF の導出
gwexpy API: Spectral module, Fitting module
関連理論: 検証済みアルゴリズム, 数値安定性
関連チュートリアル: Advanced Fitting, Noise Budget